DEFINITION
Ras proteins superfamily have served as a prototype for a group of 20-25 kDa guanine nucleotide-binding proteins that share structural homology, the superfamily of Ras-related proteins (Structure of small G proteins and their regulators, 2007). This superfamily of proteins is comprised of over 60 mammalian members and, in terms of both primary sequence and biological activity, it can be conveniently divided into several subfamilies: Ras, Rho, Rab, Arf, Ran and Rad/Gem.
THE GENE
CHEMICAL STRUCTURE AND IMAGES
When relevant for the function
- Primary structure
- Secondary structure
- Tertiary structure
- Quaternary structure
Protein Aminoacids Percentage
The Protein Aminoacids Percentage gives useful information on the local environment and the metabolic status of the cell (starvation, lack of essential AA, hypoxia)
Protein Aminoacids Percentage (Width 700 px)

SYNTHESIS AND TURNOVER
mRNA synthesis
protein synthesis
post-translational modifications
degradation
CELLULAR FUNCTIONS
cellular localization,
biological function
- Cell signaling and Ligand transport
- Structural proteins
REGULATION
DIAGNOSTIC USE
"WikiGenes":
GeneCards
iHop
OMIM
- Ras family consists of
- Ras proteins (H-Ras, K-RasA, K-RasB and N-Ras),
- four Rap proteins (Rap1A, Rap1B, Rap2A and Rap2B),
- three R-Ras-like proteins (R-Ras, TC21 and R-Ras3),
- two Ral proteins (RalA and RalB),
- the newly identified Rheb and M-Ras proteins.
This family is characterized by high similarity in the effector domain, the so-called switch I region.

CHEMICAL STRUCTURE AND IMAGES

The isoforms are principally distinguished from each other by the final 23–24 amino acid stretch, the hypervariable region (HVR), where there is < 15% sequenze similarity between Ras proteins. The HVR contains all of the motifs responsible for membrane binding and trafficking of each isoform.

Full Size Figure
Protein Aminoacids Percentage

SYNTHESIS AND TURNOVER
After synthesis on cytosolic polysomes (Compartmentalized signalling: Ras proteins and
signalling nanoclusters, 2008), Ras isoforms undergo a series of posttranslational modifications to increase their membrane affinity.

The cysteine in the C-terminal CAAX motif is farnesylated before the AAX is proteolytically cleaved and the farnesyl–cysteine is carboxymethylated . The farnesyl group promotes weak interaction with the endoplasmic reticulum (ER) which is stabilized by an adjacent set of motifs that vary among Ras isoforms. This consists of mono [N-, K(A)-Ras] or di-palmitoylation (H-Ras) of cysteines or a hexalysine polybasic sequence [K(B)-Ras].The second signal motif and farnesylated cysteineshared by all Ras isoforms comprise the targeting domain; a minimal motif that when fused to GFP displays a superficially equivalent localization as the cognate full length H- and K(B)-Ras proteins. Recent data revealed that a third signal motif is necessary for the correct localization of mono-palmitoylated N- and K(A)-Ras isoforms. GFP conjugated to the minimal targeting domain of N-Ras is restricted to the Golgi, whereas when the adjacent linker region of the HVR is included the construct localizes to the cell surface. When the HVR of the palmitoylated Ras isoforms is compared, there is 70% sequence homology including a six-residue basic ⁄ hydrophobic patch at the N-terminus of the HVR. Mutating this sequence increases the amount of endomembranous localization observed, indicating that this motif contributes to cell-surface localization. Similarly, for K(A)-Ras, the basic patch adjacent to the palmitoyl group is sufficient to ensure cell-surface localization. The second signal motif also determines the trafficking routes taken by H-, N- and K(B)-Ras to the plasma membrane. K(B)-Ras traffics via a poorly characterized Golgi-independent route that in yeast requires class C vps proteins which are normally required to regulate endosome fusion. Unlike other small G proteins such as Rabs and Rho proteins, there has been no chaperone such as GDI characterized for cytosolic Ras trafficking. Although palmitoylated Ras isoforms have also been characterized to traffic via Golgi-independent routes in yeast and adipocytes, in fibroblasts they traffic through the conventional secretory pathway.

The final destination of the post-translationally-modified GTPases depends on the computation
by cells of other ancillary signals present in the GTPase C terminus. In the case of
palmitoylated GTPases, one of these additional signals is the nature of the isoprenyl group
attached to the CAAX box.
Mapping the nucleotide and isoform dependent structural and dynamical features of Ras proteins, 2009 The Ras superfamily at a glance, 2005
Ras Isoform-Specific Signaling: Location,2001)

Like other G proteins, Ras cycles between the GDP-bound inactive form and the GTP-bound active form(Blocking Oncogenic Ras Signaling for Cancer Therapy, 2001).

CELLULAR FUNCTIONS
Ras GTPases—H-Ras, N-Ras, and K-Ras 4B/4A—operate as key molecular switches that convey extracellular signals (Blocking Oncogenic Ras Signaling for Cancer Therapy, 2001) from surface receptors to the interior of the cell, thereby regulating essential processes including proliferation, differentiation, and survival . It is well known that Ras must be attached to the inner leaflet of the plasma membrane protein (PM) ( Na+/K+-ATPase-mediated signal transduction and Na+/K+-ATPase regulation, 2008) to be functional.
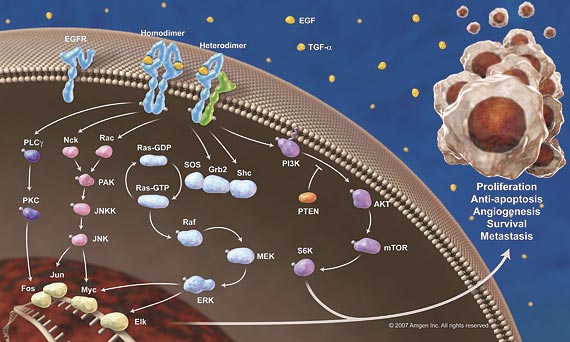
In many cell types, the MAPKs ERK1/2 are linked to cell proliferation (The roles of MAPKs in disease, 2008). ERK1/2 are thought to play a role in some cancers, because mutations in Ras and B-Raf, which can activate the ERK1/2 cascade.In human cancer ( Blocking of p53-Snail Binding, Promoted by Oncogenic K-Ras, Recovers p53 Expression and Function, 2009), the oncogenic mutation of Ras family genes including H-, N-, and K-Ras, is frequently detected in particular, K-Ras mutation is frequent event in
The major alteration known interests the methylation (DNA methylation-mediated nucleosome dynamics and oncogenic Ras signaling, 2008)and on other levels of ras pathway that involve in some somatic gene mutations (Combinatorial patterns of somatic gene mutations in cancer ,2008).
REGULATION
Ras biological activity is controlled by a regulated GDP/GTP cycle. The intrinsic GDP/GTP exchange and GTP hydrolytic activity of Ras is very low and hence, cellular control of GDP/ GTP cycling modulated by two types of regulatory proteins. Guanine nucleotide exchange factors (GEFs; RasGRF/mCDC25, SOS1/2) promote formation of the active GTP-bound state and Ras GTPase activating proteins (GAPs; p120 GAP, NF1-GAP/neurofibromin) promote formation of the inactive GDP-bound state (Increasing complexity of Ras signaling, 1998)

DIAGNOSTIC USE
Naturally as you immagine the ras pathway rappresents a “molecular target”. This is important expecially in the cancer therapies infact the decision to use some drugs are based on the activation or not of the single pathway. (A gene expression predictor of response to EGFR-targeted therapy stratifies progression-free survival to cetuximab in KRAS wild-type metastatic colorectal cancer, 2009,, Novel Agents in CML Therapy: Tyrosine Kinase Inhibitors and Beyond, 2008)