Figures heme transport
Figures progesterone+affect+hepcidin+that+regulates+ferroportin
Heme and diet
Heme constitutes 95% of functional iron in the human body, as well as two-thirds of the average person’s iron intake in developed countries. Hence, a wide range of epidemiological studies have focused on examining the association of dietary heme intake, mainly from red meat, with the risks of common diseases. High heme intake is associated with increased risk of several cancers, including colorectal cancer, pancreatic cancer and lung cancer. Likewise, the evidence for increased risks of type-2 diabetes and coronary heart disease associated with high heme intake is compelling. Furthermore, recent comparative metabolic and molecular studies of lung cancer cells showed that cancer cells require increased intracellular heme biosynthesis and uptake to meet the increased demand for oxygen-utilizing hemoproteins. Increased levels of hemoproteins in turn lead to intensified oxygen consumption and cellular energy generation, thereby fueling cancer cell progression. Together, both epidemiological and molecular studies support the idea that heme positively impacts cancer progression. However, it is also worth noting that heme deficiency can cause serious diseases in humans, such as anemia, porphyrias, and Alzheimer’s disease. This review attempts to summarize the latest literature in understanding the role of dietary heme intake and heme function in diverse diseases.
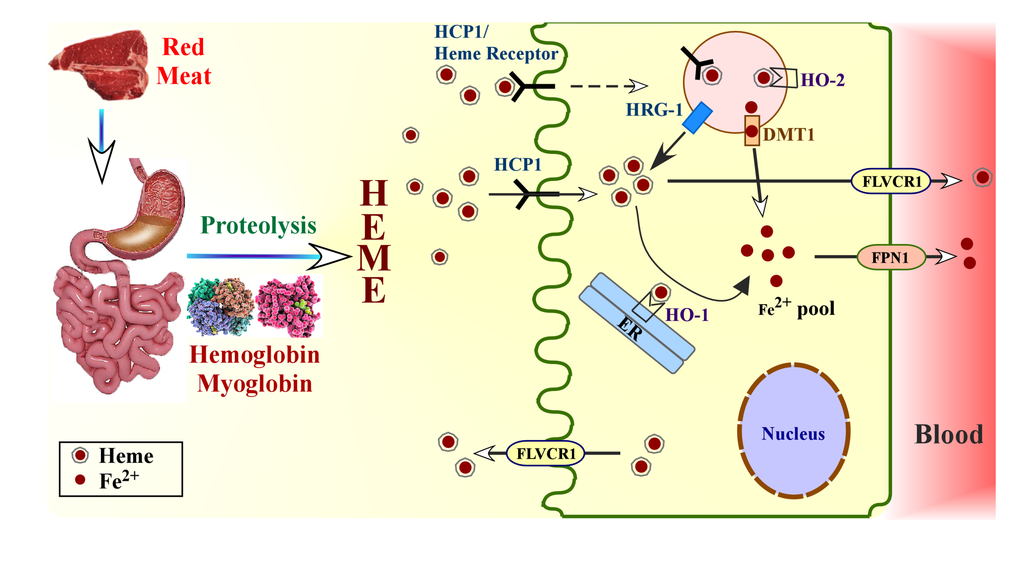
Heme absorption in gut from dietary proteins. Low pH of stomach releases heme-containing proteins hemoglobin and myoglobin from dietary meat. Heme is released by the action of proteases in stomach and intestine. Intake of heme into enterocytes can be facilitated by vesicular transport system when heme binds to heme transporter or heme receptor. Additionally, heme can be directly imported into the enterocytes by HCP1. Heme is transported to the cytoplasm from the vesicles possibly by HRG-1, and is then metabolized by HO-1 present on endoplasmic reticulum. Iron is released subsequently. Alternatively, heme inside the vesicles can be metabolized by the action of HO-2 present on vesicle membrane, and the released iron (Fe2+) is transported into the cytoplasm by metal transporter DMT1 to join the common pool of iron in the cytoplasm.
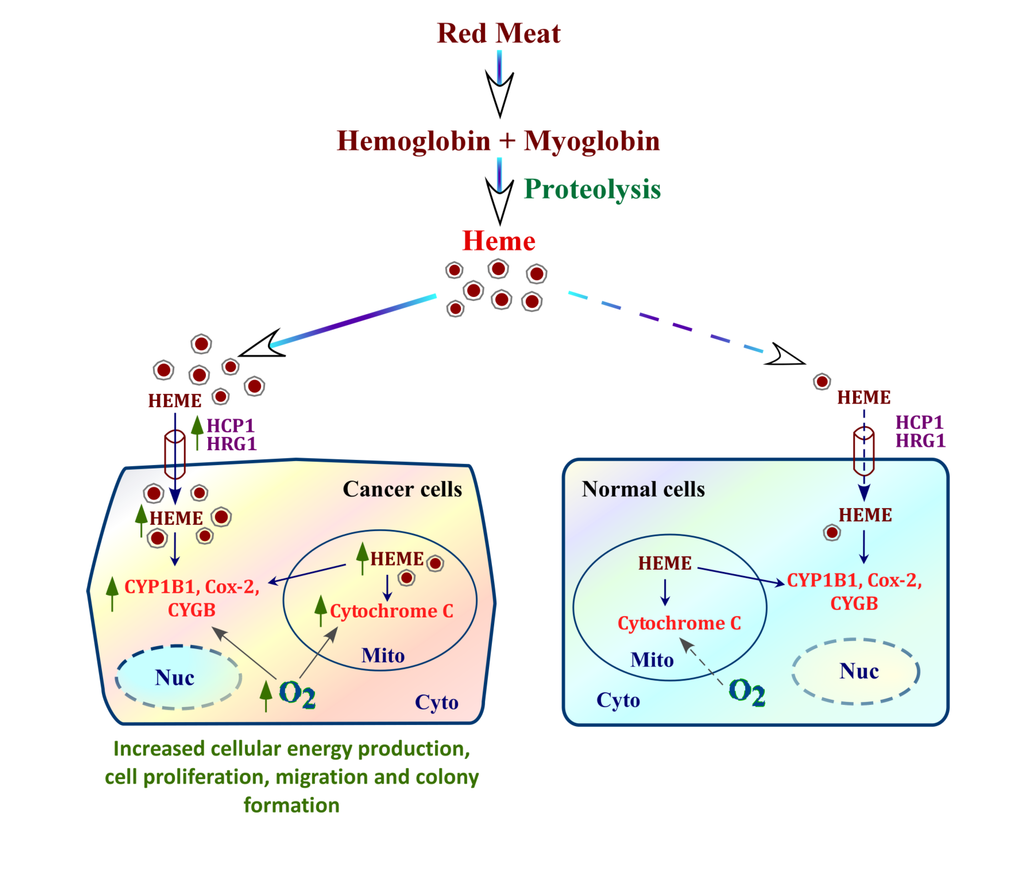
mechanism by which heme fuels cancer cell progression. Heme from blood can be taken up by cells via heme transporters HCP1 and HRG-1. Cancer cells have intensified internal heme synthesis as well as increased heme uptake via heme transporters, whose expression is dramatically elevated in cancer cells, compared to normal cells
Heme, an Essential Nutrient from Dietary Proteins, Critically Impacts Diverse Physiological and Pathological Processes, 2014

The Intestinal Heme Transporter Revealed, 2005(05)00864-0

Adult humans have about 25 trillion red blood cells (RBCs), and each second we recycle about 5 million RBCs by erythrophagocytosis (EP) in macrophages of the reticuloendothelial system. Despite the central role for EP in mammalian iron metabolism, the molecules and pathways responsible for heme trafficking during EP remain unknown. Here, we show that the mammalian homolog of HRG1, a transmembrane heme permease in C. elegans, is essential for macrophage iron homeostasis and transports heme from the phagolysosome to the cytoplasm during EP. HRG1 is strongly expressed in macrophages of the reticuloendothelial system and specifically localizes to the phagolysosomal membranes during EP. Depletion of Hrg1 in mouse macrophages causes attenuation of heme transport from the phagolysosomal compartment. Importantly, missense polymorphisms in human HRG1 are defective in heme transport. Our results reveal HRG1 as the long-sought heme transporter for heme-iron recycling in macrophages and suggest that genetic variations in HRG1 could be modifiers of human iron metabolism.
HRG1 Is Essential for Heme Transport from the Phagolysosome of Macrophages during Erythrophagocytosis
, 2013
DEFINITION
Hemopexin , binds heme with the highest affinity of any known protein. Its function of scavenging the heme released or lost by the turnover of heme proteins.
Heme scavenging protects the organism from
- heme requiring bacteria (Yersinia Pestis eg.)
- oxidative damage
THE GENE
CHEMICAL STRUCTURE AND IMAGES
When relevant for the function
- Primary structure
- Secondary structure
- Tertiary structure
- Quaternary structure
Protein Aminoacids Percentage
The Protein Aminoacids Percentage gives useful information on the local environment and the metabolic status of the cell (starvation, lack of essential AA, hypoxia)
Protein Aminoacids Percentage (Width 700 px)

SYNTHESIS AND TURNOVER
mRNA synthesis
protein synthesis
post-translational modifications
degradation
CELLULAR FUNCTIONS
cellular localization
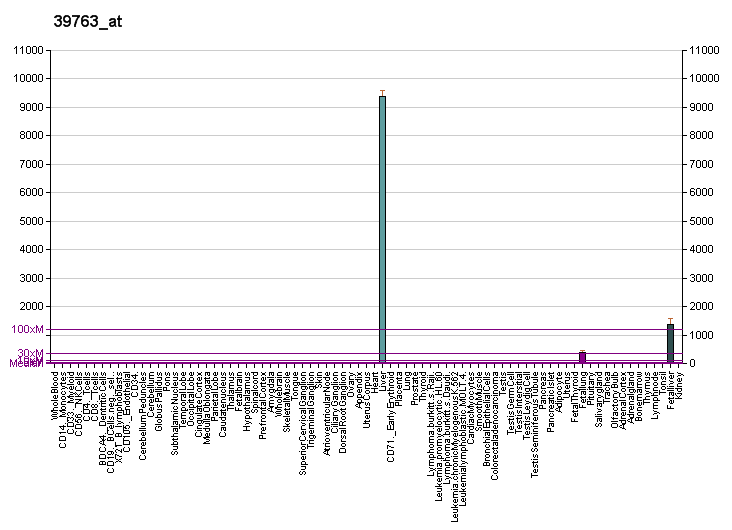
Its messenger is found only in liver as its receptor.
hepatocytes, macrophages, and syncytiotrophoblasts.
The primary receptor for uptake of Hx-heme complexes is hitherto not identified.
The asialoform of Hx has been reported to recognize the asialoglycoprotein receptor, but this hepatocyte-specific receptor is suggested to represent a secondary receptor for catabolism of outdated Hx, which has lost the terminal sialic acids of the N-linked carbohydrates
biological function
Identification of the receptor scavenging hemopexin-heme complexes 2005
Heme released from heme-binding proteins on internal hemorrhage, hemolysis, myolysis, or other cell damage is highly toxic due to oxidative and proinflammatory effects. Complex formation with hemopexin, the high-affinity heme-binding protein in plasma and cerebrospinal fluid, dampens these effects and is suggested to facilitate cellular heme metabolism. Using a ligand-affinity approach, we purified the human hemopexin-heme receptor and identified it as the low-density lipoprotein receptor-related protein (LRP)/CD91, a receptor expressed in several cell types including macrophages, hepatocytes, neurons, and syncytiotrophoblasts. Binding experiments, including Biacore analysis, showed that hemopexin-heme complex formation elicits the high receptor affinity. Uptake studies of radio-labeled hemopexin-heme complex in LRP/CD91-expressing COS cells and confocal microscopy of the cellular processing of fluorescent hemopexin-heme complex established the ability of LRP/CD91 to mediate hemopexin-heme internalization resulting in cellular heme uptake and lysosomal hemopexin degradation. Uptake of hemopexin-heme complex induced LRP/CD91-dependent heme-oxygenase 1 mRNA transcription in cultured monocytes. In conclusion, hemopexin-heme complexes are removed by a receptor-mediated pathway showing striking similarities to the CD163-mediated haptoglobin-hemoglobin clearance in macrophages. Furthermore, the data indicate a hitherto unknown role of LRP/CD91 in inflammation.
In the cell, toxic effects of heme are prevented by the activity of the heme oxygenases (HOs), which cause breakdown of the porphyrin ring into biliverdin, carbon monoxide, and iron. Biliverdin is subsequently converted to bilirubin by the biliverdin reductase, whereas iron is bound to ferritin.
- Cell signaling and Ligand transport
- Structural proteins
REGULATION
DIAGNOSTIC USE
Hemopexin and nephrotic syndrome 2008